Thursday, 2 April, 2026
Python Integration
IsoFind integrates with Python in two complementary ways: via the open-source isof package available on PyPI, which allows reading and verifying .isof files in any Python script independently of IsoFind, and via the local REST API, which allows querying and manipulating the IsoFind database directly from Python while IsoFind is open.
The isof package (open source)
PyPI Package
The isof package is an open-source reader and validator for the ISOF v1.0 format. It is independent of IsoFind: any Python script can read an .isof file without the software being installed. The source code is published under the MIT license.
pip install isof
With pandas support (recommended for data analysis):
pip install isof[pandas]
Requires Python 3.9 or higher.
Loading a file
import isof
# Load from a file
report = isof.load("bolivia_analysis.isof")
print(report)
# <ISOfDocument v1.0 — 12 sample(s) — IGE Grenoble>
# Load from a JSON string (from an API, database, etc.)
with open("analysis.isof") as f:
report = isof.loads(f.read())
Verifying integrity and authenticity
The ISOF format supports two levels of signature. The package verifies both offline, without a network connection: IsoFind Root CA and Issuing CA certificates are embedded in the package.
| Level | Mechanism | Guarantee |
|---|---|---|
| 1 | SHA-256 Hash of content | Integrity: the file has not been modified since export. |
| 2 | ECDSA P-256 Signature + IsoFind PKI | Authenticity: data was signed by an IsoFind certified laboratory. |
# Simple verification (bool)
if report.is_authentic():
print(f"Data intact. Signed by: {report.signature.signed_by}")
else:
print("Data altered or signature invalid.")
# Detailed result
result = report.verify()
print(result.valid) # bool
print(result.level) # 0 (none), 1 (SHA-256) or 2 (ECDSA PKI)
print(result.signer) # organization or CN of the certificate
print(result.signed_at) # ISO 8601 timestamp
print(result.reason) # None if valid, error message otherwise
# Distinguishing missing from corrupted signature
if result.level == 0:
print("This file is not signed.")
elif not result.valid:
print(f"Signature present but invalid: {result.reason}")
Accessing data
# Samples
print(len(report.samples))
for sample in report.samples:
print(sample.name, sample.material_type, sample.classification)
# Isotopic data for a sample
for iso in report.samples[0].isotope_data:
print(iso.element, iso.ratio_name, iso.ratio_value, iso.uncertainty)
# Document metadata
print(report.created_by.organisation)
print(report.created_at)
print(report.project)
# Methods documented in the file
for key, method in report.methods.items():
print(key, method.get("label"))
# Purification yields
for key, yield_data in report.purification.items():
print(key, yield_data) # key: "{sample_id}_{element}"
Exporting to pandas and CSV
The DataFrame produced by to_pandas() is in "tidy" format: one row per isotopic measurement, with sample metadata duplicated on each row. This format is directly compatible with matplotlib, seaborn, or any other analysis tool.
df = report.to_pandas()
# Available columns depending on file content
df[["sample_name", "element", "ratio_name", "ratio_value", "ratio_2se"]]
# Standard pandas filters
pb_data = df[df["element"] == "Pb"]
sources = df[df["classification"] == "source"]
sr_ratio = df[df["ratio_name"] == "87Sr/86Sr"]
# Quick statistics
sr_ratio["ratio_value"].describe()
# CSV Export
report.to_csv("export.csv")
# equivalent to: report.to_pandas().to_csv("export.csv", index=False)
Browsing pipeline stages
for sample in report.samples:
pipeline = report.get_pipeline(sample.id)
if pipeline:
print(f"{sample.name} — pipeline: {pipeline.label}")
for stage in pipeline.stages:
print(f" {stage.label}: {stage.status}")
Error handling
from isof.exceptions import ISOfParseError, ISOfVersionError, ISOfSignatureError
try:
report = isof.load("file.isof")
except ISOfVersionError as e:
# Format version not supported by this package version
print(f"Version {e.found} not supported — update isof.")
except ISOfParseError as e:
# Invalid JSON file or incorrect structure
print(f"Invalid file: {e}")
except ISOfSignatureError as e:
# Error during cryptographic verification
print(f"Signature error: {e}")
The package is available at pypi.org/project/isof and the source code at https://github.com/ColinFerrari/isof. The ISOF format is an open standard: third-party implementations in other languages are encouraged.
Querying IsoFind from Python via Local API
Local REST API
While IsoFind is open, its FastAPI backend is accessible at http://127.0.0.1:8001. A Python script can query this API directly using the requests library to read data, create samples, or trigger analyses from a Jupyter notebook, processing script, or external calculation pipeline.
pip install requests
Preliminary Step: Enabling API Access
By default, external API access is disabled in IsoFind. Before using a Python script, access must be enabled from the software preferences:
Preferences
→
Security
→
External API Access
Three levels are available:
| Mode | Behavior | Recommended Usage |
|---|---|---|
| Disabled | No external script can connect. /api/local/token returns 403. | Default. Keep if no external script is needed. |
| Local access without confirmation | Any script from 127.0.0.1 gets a token directly, without a popup. | Isolated personal workstation, environment of total trust. |
| Access with manual authorization | Each new program triggers a popup in IsoFind with the executable path. The user approves or refuses. "Remember" saves the program. | Shared workstation, professional environment, sensitive usage. |
In local access without confirmation mode, any malicious software installed on the workstation can read and export the entire database. Only activate this mode on a machine where you fully control the content. When in doubt, prefer the manual authorization mode.
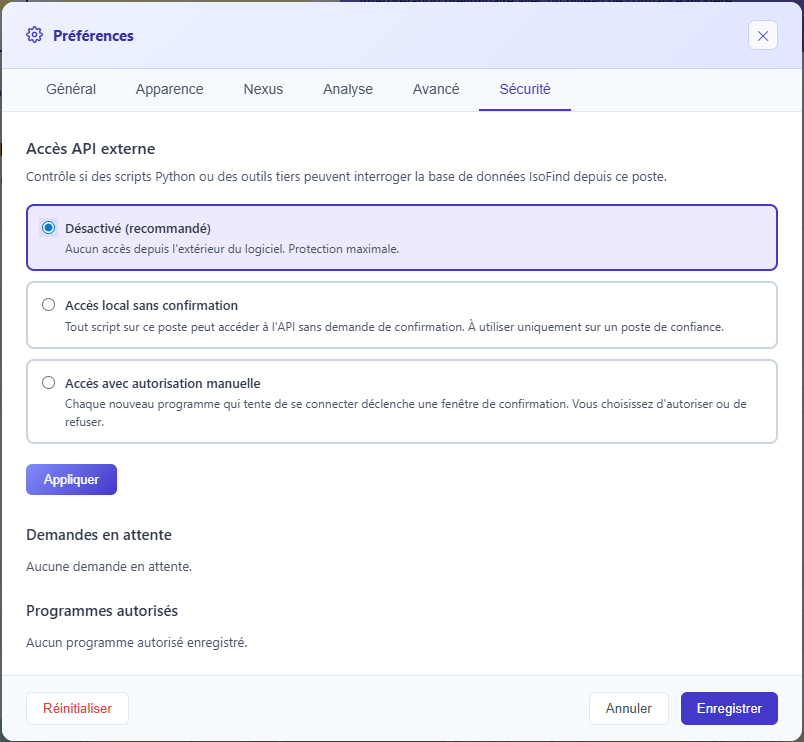 Figure 1: Security tab in preferences with API access mode selector and list of authorized programs.
Figure 1: Security tab in preferences with API access mode selector and list of authorized programs.
Obtaining the Security Token
Once access is enabled, any Python script retrieves its token via a dedicated route accessible only from 127.0.0.1. In manual authorization mode, the first call waits (up to 60 seconds) for the user to validate the popup in IsoFind.
import requests
BASE = "http://127.0.0.1:8001"
# The script sends its PID to help IsoFind identify the executable
import os
r = requests.get(
f"{BASE}/api/local/token",
headers={"X-Client-PID": str(os.getpid())}
)
# If mode is "disabled", r.status_code == 403
# If mode is "manual authorization" and user hasn't responded yet,
# the request waits up to 60 seconds
if r.status_code == 403:
err = r.json().get("error", "")
if err == "api_access_disabled":
print("API Access disabled. Enable it in IsoFind > Preferences > Security.")
elif err == "authorization_denied":
print("Access denied by user.")
raise SystemExit(1)
info = r.json()
TOKEN = info.get("token")
HEADERS = {"X-IsoFind-Token": TOKEN} if TOKEN else {}
print(f"Connected — mode: {info.get('mode', 'dev')}")
In manual authorization mode with "Remember" checked, the program is saved in IsoFind. Subsequent executions no longer display a popup: the token is returned immediately. To manage saved programs (view, revoke), go to Preferences > Security > Authorized Programs.
A malicious web browser cannot access /api/local/token: the Host header validation middleware blocks any request where the host is not 127.0.0.1:8001 or localhost:8001. A Python script on the same machine correctly sends Host: 127.0.0.1:8001 and is not affected.
Verifying that IsoFind is ready
import requests
BASE = "http://127.0.0.1:8001"
r = requests.get(f"{BASE}/api/ready",
headers=HEADERS)
print(r.json())
# {'status': 'ready'}
Retrieving samples
import requests
import pandas as pd
BASE = "http://127.0.0.1:8001"
# All samples
samples = requests.get(f"{BASE}/api/samples",
headers=HEADERS).json()
print(f"{len(samples)} samples")
# With sorting and limit
samples = requests.get(
f"{BASE}/api/samples",
params={"sort": "desc", "limit": 100}
).json()
# A specific sample
s = requests.get(f"{BASE}/api/samples/42",
headers=HEADERS).json()
print(s["name"], s["material_type"])
# Full version with methods, publications, and pipeline
s_full = requests.get(f"{BASE}/api/v2/samples/42/full",
headers=HEADERS).json()
print(s_full["methods"])
Building a DataFrame from the IsoFind database
import requests
import pandas as pd
BASE = "http://127.0.0.1:8001"
# Retrieve all isotopic data at once
iso_data = requests.get(f"{BASE}/api/samples/isotopic-data",
headers=HEADERS).json()
df = pd.DataFrame(iso_data)
# Available columns: sample_id, sample_name, element, ratio_name,
# ratio_value, uncertainty, standard_used, instrument, normalized, ...
print(df.columns.tolist())
# Statistics by element
print(df.groupby("element")["ratio_value"].describe())
# Filter for normalized samples only
df_norm = df[df["normalized"] == True]
Creating a sample
r = requests.post(
f"{BASE}/api/samples",
headers=HEADERS,
json={
"name": "BIF-2026-001",
"material_type": "rock",
"sector": "geology",
"collection_date": "2026-02-15",
"collection_location":"Cerro Rico, Bolivia",
"classification": "source",
"latitude": -19.6078,
"longitude": -65.7527,
"project": "antimony-traceability-2026",
}
)
print(r.status_code, r.json())
Recording purification yields
sample_id = 42
# Record Sb yield for this sample
r = requests.post(
f"{BASE}/api/samples/{sample_id}/purification-yields",
json={
"element": "Sb",
"yield_percent": 87.3,
"operator": "C. Ferrari",
"method_key": "ion-exchange-sb-v2"
}
)
print(r.json())
# {'success': True, 'sample_id': 42, 'element': 'SB', 'yield_percent': 87.3}
# Read recorded yields
yields = requests.get(
f"{BASE}/api/samples/{sample_id}/purification-yields",
headers=HEADERS
).json()
print(yields)
Running a matching search
# Analyze a sample already in the database
r = requests.post(
f"{BASE}/analyze",
headers=HEADERS,
json={
"sample_id": 42,
"threshold": 0.85,
"algorithm": "hybrid",
"classification_filter": "sources",
"same_material": True,
}
)
result = r.json()
print(f"{len(result['matches'])} matches found")
for match in result["matches"][[:5]]:
print(f" #{match['rank']} {match['zone_name']} — score {match['overall_score']:.3f}")
# Manual analysis without sample in database
r = requests.post(
f"{BASE}/analyze/manual",
headers=HEADERS,
json={
"sample_name": "Unknown-01",
"material_type":"ore",
"threshold": 0.75,
"isotope_data": [
{"element": "Sb", "ratio_name": "123Sb/121Sb",
"ratio_value": 0.74738, "uncertainty": 0.00012},
{"element": "Pb", "ratio_name": "206Pb/204Pb",
"ratio_value": 18.421, "uncertainty": 0.003},
]
}
)
print(r.json()["matches"][[:3]])
Batch Analysis
# Analyze multiple samples in a single request
r = requests.post(
f"{BASE}/api/matching/batch",
json=[1, 2, 3, 42, 57], headers=HEADERS
)
results = r.json()
for sample_id, res in results.items():
if res["status"] == "success":
print(f"ID {sample_id}: {res['matches_count']} matches, "
f"best zone = {res['best_zone']} (score {res['best_score']:.3f})")
else:
print(f"ID {sample_id}: error — {res['error']}")
Exporting to CSV from Python
import requests
# Download complete database CSV
r = requests.get(f"{BASE}/api/samples/export",
headers=HEADERS)
with open("export.csv", "wb") as f:
f.write(r.content)
# Grouped export with all associated data (methods, publications)
r = requests.post(
f"{BASE}/api/v2/samples/export-batch",
json=[1, 2, 3]
)
data = r.json()["data"]
print(f"{data['count']} samples exported on {data['exported_at']}")
Querying CRMs and shifts
# List available CRMs
crms = requests.get(f"{BASE}/api/crm/list",
headers=HEADERS).json()["data"]
print(f"{len(crms)} CRMs in database")
# Calculate shift for a given CRM
r = requests.post(
f"{BASE}/api/shifts/calculate/NIST-3102a",
params={
"element": "Sb",
"isotope_ratio":"123Sb/121Sb"
}
)
shift = r.json()
if shift["success"]:
print(f"Calculated shift: {shift['data']}")
# Convert an isotopic value from one standard to another
r = requests.post(
f"{BASE}/api/shifts/convert",
json={
"value": 0.74738,
"from_std": "NIST-3102a",
"to_std": "IAEA-SbS-1",
"element": "Sb",
"ratio_name": "123Sb/121Sb"
}
)
print(r.json())
Interactive Swagger documentation is available at http://127.0.0.1:8001/docs while IsoFind is open. It lists all routes with their full JSON schemas and allows testing them directly from the browser, which is useful for exploring new routes before integrating them into a script.
Combining Both Approaches
All helper functions defined below assume that BASE and HEADERS are defined at the top of the script via GET /api/local/token. In a Jupyter notebook, run the initialization cell once at the start of the session.
The most powerful workflow combines the local API (to access live IsoFind database data) and the isof package (to read and verify exported files). Full example: export a set of samples from IsoFind, sign the resulting ISOF file, and verify it programmatically.
import requests
import isof
import pandas as pd
BASE = "http://127.0.0.1:8001"
# 1. Retrieve samples from the IsoFind database
iso_data = requests.get(f"{BASE}/api/samples/isotopic-data",
headers=HEADERS).json()
df = pd.DataFrame(iso_data)
# 2. Analyze: means by element and material
pivot = df.pivot_table(
values="ratio_value",
index="sample_material_type",
columns="element",
aggfunc="mean"
)
print(pivot)
# 3. Load and verify an ISOF file exported by IsoFind
report = isof.load("certified_export.isof")
result = report.verify()
if result.valid and result.level == 2:
print(f"Certified by: {result.signer}")
print(f"Signed on: {result.signed_at}")
df_certified = report.to_pandas()
else:
raise ValueError(f"Invalid signature: {result.reason}")
Built-in Python Console in IsoFind
Pyodide Console
In addition to the two approaches above, IsoFind includes an interactive Python console that runs directly within the software, with no installation required. It is powered by Pyodide (Python 3.11 compiled to WebAssembly) and provides access to current database data via ready-to-use variables and functions.
The console is accessible from:
Tools Menu
→
Python Console
 Figure 2: IsoFind Python console with database data loaded automatically.
Figure 2: IsoFind Python console with database data loaded automatically.
Automatically Available Variables
# Loaded at startup from /api/samples
samples # list[dict]: all samples in the database
isotopes # dict: isotopic data indexed by sample name
Pre-defined Utility Functions
# Access by ID or name
s = get_sample(42)
s = get_sample_by_name("BIF-2026-001")
iso = get_isotopes("BIF-2026-001")
# Convert to pandas DataFrame (one row per isotopic measurement)
df = to_dataframe()
print(df.head())
# Statistics for an isotope ratio
isotope_stats("87Sr/86Sr")
Statistics for 87Sr/86Sr:
Count: 42
Mean: 0.710341
Std: 0.002187
Min: 0.706012
Max: 0.715892
Median: 0.710215
Available Libraries
import numpy as np
import pandas as pd
# Example: distribution of 87Sr/86Sr values
vals = [
iso.get("ratio_value")
for s in samples
for iso in (s.get("isotope_data") or [])
if iso.get("ratio_name") == "87Sr/86Sr"
and iso.get("ratio_value") is not None
]
arr = np.array(vals, dtype=float)
print(f"n={len(arr)}, mean={arr.mean():.6f}, std={arr.std():.6f}")
# Example: filtering and sorting with pandas
df = to_dataframe()
rocks = df[df["material_type"] == "rock"].sort_values("87Sr/86Sr")
print(rocks[["name", "87Sr/86Sr", "87Sr/86Sr_err"]])
Matplotlib is not available in the Pyodide console (no graphical rendering in the browser environment). For visualizations, use df.describe() and isotope_stats() for statistics, or export the data and plot it in an external Python script.
Summary of the Three Approaches
| Approach | When to use it | IsoFind Required | Installation |
|---|---|---|---|
| isof package (PyPI) | Read and verify .isof files in any script or pipeline, independently of IsoFind. Share verifiable data with third parties. | No | pip install isof |
| Local API (requests) | Access the live IsoFind database, create samples, trigger analyses from a Jupyter notebook or external processing script. Requires enabling access in Preferences > Security. | Yes (Running) | pip install requests |
| Built-in Pyodide Console | Quick exploration, ad hoc calculations, statistics on the current database. No installation, directly inside IsoFind. | Yes (Running) | None |
